
This is another parameter that can be used to adjust primer specificity stringecy. Try to lower the mismatch value in such case. However, specifying a larger mismatch value may make it more difficult to find such specific primers. The larger the mismatches (especially those toward 3' end) are between primers and the unintended targets, the more specific the primer pair is to your template (i.e., it will be more difficult to anneal to unintended targets). This requires at least one primer (for a given primer pair) to have the specified number of mismatches to unintended targets. You can use your own sequences (accession number, gi, or FASTA sequence) as a search database. This database is recommended if you are not considering variations represented by alternate loci. Mitochondrion and plastid genomes are also included where applicable.Īlthough sequences in this database are completely covered by the Refseq representative genomes database, it does not contain the alternate loci and thus avoids sequence redundancy introduced by including alternate loci. These are Refseq representative genomes from primary chromosome assemblies (i.e., no alternate loci) for many eukaryotic organisms. Genomes for selected eukaryotic organisms (primary assemblies only): This contains all RNA entries from NCBI's Reference Sequence collection 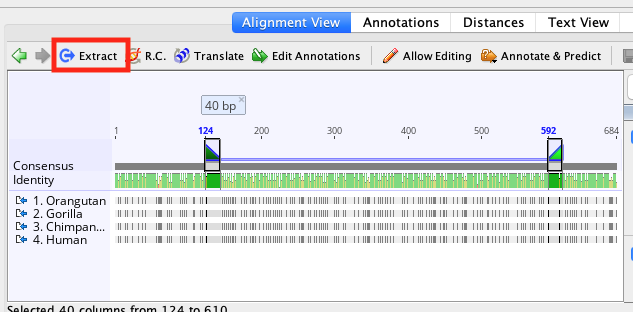
Mitochondrion genomes are included where applicable. For other species, genomes from diverse isolates of the same species may be included. For the eukaryotes, only one genome is included per species (However, alternate loci of eukaryotic genomes are included where applicable). This database contains minimum redundancy in genome representation. These genomes are among the best quality genomes available at NCBI. This database contains NCBI RefSeq Reference and Representative genomes across broad taxonomy groups including eukaryotes, bacteria, archaea, viruses and viroids. You will then be given the option to delete the pair of that primer at the same time.This contains mRNA only from NCBI's Reference Sequence collection To delete primers that you don’t want, just select the primer annotation and click the Delete button. 
In the case of the reverse primer it will automatically be reverse complemented. This will generate a separate, short sequence document in oligo format which just contains the primer sequence and the annotation (which contains the primer characteristics). The best way to save a primer or DNA probe for further testing or use is to select the annotation for that primer and click the Extract button in the sequence viewer. Mismatches between the primer and off-target will be shown in red. An off-target site will be listed if it has no mismatches to the first four 3 ′ bases of the primer, and less than 10% mismatches with the primer overall. The entire sequence will be searched for off-target sites, even if only a selected region is chosen for primer design. These are putative non-specific primer binding sites identified on the sequence that was used for primer design. In Geneious Prime 2019.1 onwards, the primer annotation includes a list of Off-target sites for that primer, including their location and sequence. All other values including hairpin and self dimer T m are calculated with extension included. Note that in Geneious Prime 2020 onwards, for primers with 5 ′ extensions the primer length, T m and %GC is calculated both with and without the extension. Shows how the values in the Geneious primer annotation map to the original Primer3 values. Alternatively, double clicking on an annotation will display its details in the annotation editing dialog. The information will be presented in a popup box. Primers will be coloured green and probes red.ĭetailed information such as melting point, tendency to form primer-dimers and GC content can be seen by hovering the mouse over the primer annotation. The annotations will be labelled with the base number the primer starts at, followed by either F (forward primer), R (reverse primer), or P (probe). When complete, primers and probes will be added as annotations on the sequences. A progress bar may appear for a short time while the process completes. Once the task and options have been set, click the OK button to design the primers.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
